Involvement of multiple P450s and UDP-GTs in the in vitro metabolism of Muraglitazar
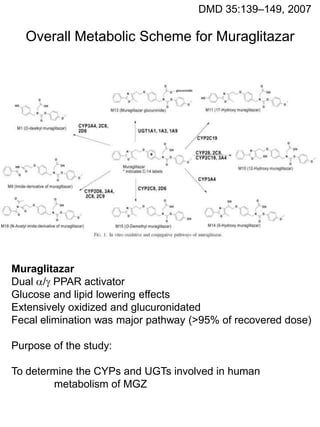
- 1. DMD 35:139–149, 2007 Overall Metabolic Scheme for Muraglitazar Muraglitazar Dual / PPAR activator Glucose and lipid lowering effects Extensively oxidized and glucuronidated Fecal elimination was major pathway (>95% of recovered dose) Purpose of the study: To determine the CYPs and UGTs involved in human metabolism of MGZ
- 2. Oxidation of Muraglitazar in Humans minor
- 3. Muraglitizar Metabolism (in vivo)
- 5. Materials • Radiolabled muraglitazar (MGZ, 99.6% pure, 8.4 µCi/mg) • CYP Inhibitors: – Furafylline (1A2), 8-MOP (2A6), orphenadrine (2B6), sulfaphenazole (2C9), tranylcypromine (2C19), quinidine (2D6), ketoconazole (3A4), montekulast (2C8), benzylnirvanol (2C19), 1-ABT (general CYP) • Baculosomes of: – CYPs 1A2, 2A6, 2B6, 2C9, 2C18, 2C19, 2D6, 2E1, 3A4 and 3A5 – UGTs 1A1, 1A3, 1A4, 1A6, 1A8, 1A9, 1A10, 2B4, 2B7, and 2B15 • Monoclonal Antibodies for: – CYPs 1A2, 2A6, 2B6, 2C9, 2Cs, 2D6, 2E1, and 3A4
- 6. HLMs • A: – 360 pmol/mg CYP – 320 pmol/mg CYP reductase – 570 pmol/mg cytochrome b5 – Activities of probe substrates • CYP2C8 – 240 pmol/min/mg (paclitaxel) • CYP2C9 – 3100 pmol/min/mg (diclofenac) • CYP2C19 – 48 pmol/min/mg (S-mephenytoin) • CYP2D6 – 71 pmol/min/mg (bufuralol) • CYP3A4 – 6800 pmol/min/mg (testosterone) • B: – 240 pmol/mg CYP ( 70%) – 340 pmol/mg CYP reductase ( 100%) – 500 pmol/mg cytochrome b5 ( 90%) – Activities of probe substrates • CYP2C8 – 500 pmol/min/mg ( 200%) • CYP2C9 – 1700 pmol/min/mg ( 40%) • CYP2C19 – 26 pmol/min/mg ( 40%) • CYP2D6 – 70 pmol/min/mg ( 100%) • CYP3A4 – 4200 pmol/min/mg ( 60%)
- 7. Incubations • Run in duplicate • 14C-MGZ(26.6 µM, 25 µM, or 0-150 µM) • CYP Baculosomes (40 or 200 pmol/mL) or – HLM-A (1 mg/mL) or – baculosomes + heat inactivated HLM-A (1 mg/mL) – Human hepatocytes (2 X 106 cells/mL) • 1 mM NADPH • 100 mM NaPO4 (pH 7.4) • 0.25 mL or – 2 mL (hepatocyte studies) • 20 or 30 min at 37 C with shaking at 100 rpm or – 3 hr at 37 C (hepatocyte studies) • Kill w/ 0.25 mL ACN or – 2 mL (hepatocyte studies) • Centrifuge 10 min at 2000g • Inject 75 µL or 100 µL onto HPLC
- 8. Kinetic Parameters for MGZ glucuronidation by UGTs
- 9. ID of major MGZ metabolites by MS/MS
- 10. Inhibition of MGZ depletion • 14C-MGZ (2.5 µM) • HLM-B (0.5 mg/mL) • Chemical and Antibody Inhibitors – Montelukast (3 µM) – Sulfaphenazole (10 or 20 µM) – Benzylnirvanol (1 µM) – Tranylcypromine (30 µM) – Quinidine (1 or 15 µM) – Ketoconazole (1 or 10 µM) – 1-ABT (1000 µM) – Anti-CYP monocolonal antibodies (2-5 µL) • 1.2 mM NADPH • 50 mM NaPO4 (pH 7.4) • 5 mM MgCl2 • 0.2 mL • 30 min at 37 C with shaking at 100 rpm • Kill w/ 0.4 mL ACN containing 3% AA and 1.5 µg/mL IS • Centrifuge 15 min at 2000g • Dilute supernatant with ACN/water (2:1, v/v) • Analyze by LC/MS
- 11. Inhibition of MGZ metabolite formation • 14C-MGZ (5.17 mM) • HLM-A (1 mg/mL) • Chemical Inhibitors • 1 mM NADPH • 100 mM NaPO4 (pH 7.4) • 5 mM MgCl2 • 1 mL • 15 min at 37 C with shaking at 100 rpm or – Preincubate with MBIs + NADPH for 10 min • Kill w/ 1.5 mL ACN • Centrifuge 15 min at 2000g • Dry supernatant under N2 • Reconstitute in 300 µL water ACN (2:1, v/v) • Inject 75 µL on HPLC
- 12. UGT Incubations with HLM-A • Run in triplicate for kinetic studies • 14C-MGZ (4 µM, 26.6 µM or 0-30 µM) • HLM-B (0.8 mg/mL) or – UGT enzyme (0.1 mg/mL) • 25 µg/mL alamethicin • 25 mM Tris-HCl (pH 7.5) • 10 mM MgCl2 • 0.25 mL • Preinubate at 37 C for 5 min with shaking at 100 rpm • Initiate reaction with UDPGA (2.5 mM) • 30 min at 37 C • Kill w/ 0.5 mL ice-cold ACN • Centrifuge 15 min at 2000g • Inject 100-200 µL onto HPLC
- 13. HPLC • Radioactivity Profiling – Shimadzu Class VP system – Diode array detector (SPD-M10A) – YMC ODS AQ C-18 (5 µm, 4.6 X 150 mm, Waters) – 1 mL/min – 96-well LumaPlates used to collect 0.26 min fractions – Gilson model 202 fraction collector – 70 min run – Dry plates on a SpeedVac – Analyze on a TopCount – Solvent A = 0.06% TFA, Solvent B = ACN + 0.06% TFA – Subtract out the average cpm value from first 8 samples – Total amount of each metabolite was calculated based on the percentage distribution and the total amount of parent compound used in the incubation
- 14. Metabolic Profile of MGZ LC/radiomatic analysis 26.6 µM MGZ 1 mg/mL HLM-A 30 min Phase 1 4 µM MGZ Alamethicin 30 min Phase 2 25 µM MGZ 3 hr Phase 1 + 2 16 oxidative metabs In vivo
- 16. Metabolic profile of MGZ by CYPs 26.6 µM MGZ 200 pmol/mL CYP 30 min Parent drug is the Major peak M10, M11, M14, M15 were very minor Metabolites (<0.5%)
- 17. Metabolic Profiles of MGZ by CYPs CYPs involved 2C8, 2C9, 2C19, 2D6, 3A4 CYPs not involved: 1A2, 2A6, 2B6, 2C18, 2E1, 3A5
- 18. Inhibition of MGZ depletion by MAbs and chemicals 2.5 µM MGZ (plasma concentration) 0.5 mg/mL HLM-B Inhibitor 30 min (5 µM) (montelukast, 3 µM) (sulfaphenazole, 10 µM) (benzylnirvanol, 1 µM ) (21.1%) (quinidine, 1 µM) (21.4%) (ketoconazole, 1 µM) (39.9%) (1-ABT, 1000 µM) mABs of CYPs 1A2, 2A6, 2B6, and 2E1 inhibited < 10%
- 19. Metabolic Profile of MGZ in HLM-B Identify which metab CYP 2C8 (approx 38% of total) forms HLM-B 25 µM MGZ 30 min Formation of M15: HLM-A << HLM-B 2X more 2C8 Activity in HLM-B (2C8 inhibitor)
- 20. Metabolic Profile of MGZ Compare to HLM-B
- 21. Inhibition of MGZ metabolism in HLMs 25 or 26.6 µM MGZ Inhibitors of CYPs 1A2 and 2B6 showed minor inhibition Inhibitor of CYP 2A6 showed minor activation pure CYP HLM-B CYP2C8 M10, M15 M10, M15 CYP2C9 M10 M10, M11, M14 CYP2C19 M10, M11 M10, M11, M15, M15 CYP2D6 M10 M11, M14 CYP3A4 M10, M14 M10, M11, M14, M15
- 22. Km/Vmax determinations for MGZ oxidation by CYPs 0-150 µM MGZ 40 pmol CYP 30 min
- 23. Kinetic Parameters for MGZ oxidation by CYPs Michaelis-Menten analysis Formation mg/mic Relative of each protein contribution metab by of each CYP each CYP [CYPs] in mics (pmol/mg): 2C8: 64 2C9: 96 2C19: 19 2D6: 10 3A4: 108
- 24. Possible Problems with the Calculations Vs = A3A4v3A4 + A2C9v2C9 + A2D6v2D6 + ….. A = relative abundance of each CYP in HLMs Vmax for isoform specific rxn in HLMs RAF = -------------------------------------------------------------------- Vmax for isoform specific rxn by pure CYP system (1) RAF was from a group at Merck (2) Assumes the same amount of uncoupling with every substrate/CYP combination (3) Used different isoform-selective substrates (4) Used different microsomes different levels of CYP reductase, CYP, b5 polymorphisms (4) Used different pure CYP systems lymphoblasts vs. baculosomes
- 25. Relative contributions of CYPs to MGZ oxidation Values for overall metabolism don’t change much from 1-25 µM CYP2Cs account for 60% CYP2D6 accounts for < 1% CYP3A4 account for ~ 40%
- 26. Overall Metabolic Scheme for MGZ Coadministration of ketoconazole (CYP3A4 inhibitor) or gemfibrozil (CYP2C and UGT1A1 inhibitor) did not affect the clearance of muraglitazar
- 27. Glucuronidation of MGZ HLM-A or UGT 26.6 µM 30 min The MGZ acyl glucuronide was stable at RT if the sample was acidified immediately and stored at -20°C Next: kinetic analysis (0-30 µM MGZ) Km 3 µM in HLMs and for the 3 UGTs Cannot estimate relative contributions since the [UGTs] in the liver are not known
- 28. Summary/Conclusions • MGZ is not extensively metabolized in vitro (16%) by CYPs but is metabolized at several site by several different CYPs. • Selective inhibitors and mAbs were used to support the results from metabolism experiments using individual CYPs. • MGZ appears to be glucuronidated by UGTs (glucuronides account for 80% of radioactivity in vivo). • The extensive metabolism observed for MGZ should lead to similar clearance among patients since >> one enzyme is involved in its metabolism. • The results from the individual CYP and selective CYP inhibition do not seem to match up. Might be due to submaximal inhibition of the compounds, non-specific inhibition. • Possible calculation issues.
