Final CSPG4 Presentation Abhinav Bhaskar 4-22-15.pptx
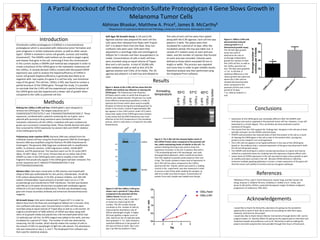
- 1. A Partial Knockout of the Chondroitin Sulfate Proteoglycan 4 Gene Slows Growth in Melanoma Tumor Cells Abhinav Bhaskar, Matthew A. Price§, James B. McCarthy§ §Department of Laboratory Medicine and Pathology, University of Minnesota, Minneapolis 55455 Introduction Methods Conclusions References Results Making the 1205Lu 2.4F6 cell line: CRISPR gRNA’s were designed to remove the CSPG4 gene. The targets sequences are 5’ CGAGCGCGGCTCTGCTCCTG 3’ and 5’AGAGACCTGGAGACACCAGG 3’. These sequences, combined with a plasmid containing the cas 9 gene and a plasmid with puromycin drug resistance were transfected into the metastatic melanoma cell line 1205Lu. Individual cells were isolated and grown up in puromycin containing media. These clonally derived cells were then screened for CSPG4 expression by western blot and CRISPR deletion of the CSPG4 gene by PCR. Polymerase chain reaction (PCR): Genomic DNA was isolated from the individual cloned cell lines using the PureLink genomic DNA kit (Invitrogen). Genomic DNA was amplified using the Platinum Pfx DNA Polymerase kit (Invitrogen). The genomic DNA (1μg) was combined with 1x amplification buffer, 1x enhancer solution, 1mM magnesium sulfate, 10mM dNTP mixture, and Pfx polymerase. The reactions were run for 35 cycles (94oC 30sec, 55-60oC 30sec, 72oC 1min). Primers that are on either side of the CRISPR cut sites in the CSPG4 gene were used to amplify a short DNA fragment that would only appear if the CSPG4 gene had been removed. The primer sequences are 5’ GGGCCCTTTAAGAAGGTTGA 3’ and GTTTTGACAGCCCAAACCAG. Western Blot: Cells were rinsed with 1x PBS solution and treated with 250μl of RIPA lysis buffer(50mM Tris HCL pH 8.0, 150mM NaCl, 1% NP-40, 0.5% sodium deoxychlorate, 0.1% SDS, protease inhibitor, and 100 mM sodium orthovandate). Equal amounts of protein were run on a 7.5% acrylamide gel and transferred to PVDF membrane. The blot was blocked with PBS w/ 0.1% tween-20 and then incubated with antibodies against CSPG4 (9.2.27) and Tubulin (Calbiochem). The blot was developed using goat-anti-mouse secondary antibody and enhanced chemoluminescence reagents. 2D Growth Assay: Cells were released with Trypsin EDT-1 in order to detach them from the flask and centrifuged at 500rpm for 5 minutes. Only non-confluent cells were used. Concentrations in both cell lines were recorded using an equal volume of Trypan Blue as well as a cell counter. An original concentration of 5000 cells/mL of CSPG4 was taken along with 10mL of 2x growth media and placed into a 96 microwell plate which had 12 replicates per cell line. An MTS reagent was added to the wells, and was incubated for a period of 7 days. The number of cells was observed by measuring the OD number, which directly relates the number of cells to the amount of 495 nm wavelength of light the cells absorb. The absorbance rate was measured on day 2, 5, and 7. The Graphpad Prism software was then used for statistical analysis. Figure 4: 1205Lu 2.4F6 cells demonstrated reduced colony growth in a 3 dimensional growth assay.. The 3D Soft Agar growth assay was used to determine the rate of anchorage-independent growth for colonies in both the 2.4F6 cell line, as well as the 1205Lu parental cell line. The data were graphed +/- S.E. In all trials. A statistical difference in the colony growth was observed where the 2.4F6 cell line showed decreased growth when compared to the parental cell line over a time period of 14 days. *=p <.005 by student’s t- test. Figure 3: Cell line 1205Lu 2.4F6 grew slower over a period of 7 days when compared to the parental cell line 1205Lu Parent. Results were measured on day 2, day 5, and day 7 as shown by measuring the OD number. The OD number directly correlates to the number of cells by measuring the absorbance rate of 495nm wavelength of light. A higher OD level signifies a higher count of cells. Data from the 12 replicate wells were graphed +/- S.E. Data showed a statistical difference in the growth of the two cell lines on both day 5 and day 7 (p<.001 by student’s t-test). Chondroitin sulfate proteoglycan 4 (CSPG4) is a transmembrane proteoglycan which is associated with melanoma tumor formation and poor prognosis in certain melanoma strains, as well as other cancer types1. CSPG4 is involved in tumor cell growth, survival, and motility (movement)1. The CRISPR–cas9 method can be used to target a gene and cleaves that gene in the cell, removing it from the chromosome1. In the current studies a CRISPR-cas9 method was employed in order to create a knockout of the CSPG4 gene in the metastatic melanoma cell line 1205Lu. A clonally derived 1205Lu mutant with decreased CSPG4 expression was used to analyze the haploinsufficiency of CSPG4 in tumor cell growth (haploinsufficiency is generally described as an organism with two copies of a gene in a cell has only one functional copy of the gene). The cell line, 1205Lu 2.4F6, was found to contain a partial knockout of the CSPG4 gene. Through the study, it is possible to conclude that the 2.4F6 cell line experienced a partial knockout of the CSPG4 gene and also experienced a slower rate of growth when compared to the 1205 Lu parental cell line. 1Matthew A Price, Leah E Colvin Wanshura, Jianbo Yang, Jennifer Carlson, Bo Xiang, Guiyuan Li, Soldano Ferrone, Arkadiusz Z. Dudek, Eva A. Turley, and James B. McCarthy, CSPG4, a potential therapeutic target, facilitates malignant progression of melanoma. PMC 2012. • Expression of the CSPG4 gene was noticeably different after the CRISPR-cas9 technique was used as opposed to the parental tumor cell line. However, it was still expressed to a certain degree. Thus, we believe we have produced a partial knockout of the gene. • The results from the PCR support the findings that the gene in the cell was at least partially removed via the CRISPR-cas9 technique. • CRISPR-cas9 procedure has reduced levels of both the protein in the cell as a result of cleaving the CSPG4 gene in the cell as seen in the western blot, causing less proteins to be created in turn and expressed. • The 2.4F6 cell line appears to be haploinsufficient in the case of the CSPG4 gene based on the evidence that a reduced expression of the gene was observed in both the 2D and 3D growth assays. • The CRISPR-cas9 technique is useful in producing knockouts as a gene and may be utilized as a potential means for studying how the expression of genes affects cells. • Lower levels of CSPG4 expression in the cell may lead to lower tumor growth as well as motility and lower survival in the cell. Because CSPG4 directly or indirectly enhances multiple signaling pathways in tumors, a lower expression of this gene will limit the tumor cell’s ability to function and use oncogenic pathways. Acknowledgements Figure 2: The 2.4F6 cell line showed higher levels of the CSPG4 Protein when compared to the parental cell line, while maintaining levels of tubulin in the cell. The western blotting technique was used to show the expression of protein in the cell. A western blot with a 7.5% acrylamide gel and a 4% stacking gel was used during gel electrophoresis, which moved the proteins from the negative to positive poles based on their size in bps. The results showed a lower level of expression in the 2.4F6 cell line when compared to the 1205Lu parental cell line. Tubulin, which was used as a loading control in this experiment, was also measured in order to ensure a lack of bias when loading the samples, as well as to make sure that an equal concentration of protein from each sample was loaded onto the gel . * Figure 1: Bands of the 2.4F6 cell line show that the CRISPR-cas9 method was effective in cleaving the CSPG4 gene. The Polymerase chain Reaction (PCR)was used in order to verify that the gene of interest was indeed cleaved. The gel was run under three different temperature gradients in order to optimize the Primers which were used to amplify the gene of interest during the annealing period. As shown in the results, a band of approximately 300 bps appears in the 2.4F6 column under 55oC which shows that copies of the CSPG4 gene were cleaved. It also shows that the DNA Polymerase was most effective at the 55oC temperature in the annealing process, which is why there is no band for the other temperatures. Soft Agar 3D Growth Assay: A 1% and 0.3% Agarose solution was prepared for each cell line. Cells were then released from flasks with Trypsin EDT-1 to detach them from the flask. Only non- confluent cells were used. Cells were then aliquoted to a centrifuge tube and centrifuged at 500rpm for 5 minutes and then resuspended in 2x media. Concentrations of cells in both cell lines were recorded using an equal volume of Trypan Blue and a cell counter. A total of 10,000 cells were needed per well as well as 4mL of 0.3% agarose solution and 3.9mL of 2x media. The 1% agarose was plated in a 6 well tray and allowed to cool. The cells of each cell line were then plated along with the 0.3% agarose. Each cell line was plated in 3 wells. The plates were then incubated for a period of 14 days. After the incubation period, the tray was taken out. A sample of 5 random areas on each well was taken, and the number of colonies that formed on each area was recorded. Colonies were defined as those which exceeded 50 mm in length or width. This process was repeated one more time in order to gain reliable results. Statistical analysis was then performed using the Graphpad Prism software. I would like to thank the McCarthy Laboratory for giving me the wonderful opportunity to work in their lab as well as for providing me with their space, materials, and time for this project. I would also like to thank Honors Mentor Connections through district 287, and its program leader, Dr. Dorothy Welch for giving me the opportunity to interview such iimportant people and professors such as Dr. McCarthy as well as making this project and other projects like mine possible through their hours of hard work.