Denunciar
Compartilhar
Baixar para ler offline
Recomendados
The presence of circulating (cell-free) nucleic acids in the bloodstream offers a potential non-invasive approach to monitor disease status and guide treatment options. In past years, increasing interest has been shown for circulating RNA; especially circulating small RNAs for their potential application as biomarkers that may lead toward more effective diagnosis and prognosis in the future. However, widespread inconsistencies have been observed among the studies due to biases generated during sample collection, handling, RNA extraction and analysis. We have developed a complete workflow that includes blood collection, plasma preparation, circulating RNA extraction, followed by expression analysis and gene fusion detection on Ion TorrentTM Next-Generation Sequencing platforms. Blood plasma research samples from normal research samples were utilized for circulating RNA isolation following a TRIzolTM LS Reagent and mirVanaTM miRNA Isolation Kit-based method to maximize circulating RNA recovery. Ion AmpliSeqTM library preparation was performed on purified circulating RNA using either Ion AmpliSeq Transcriptome panel for expression profiling of 21K coding and non-coding genes, or an Ion AmpliSeq panel targeting fusion transcript detection from RNA. Ion AmpliSeq Transcriptome data was analyzed using ampliSeqRNA plugin in Torrent Suite™ Software. ~3000 genes were detected in cfDNA from plasma research samples with high correlation (r>0.8) observed between normal research samples. Ion Reporter™ Software was used to analyze fusion transcript panel data. Detection of fusion gene transcripts was demonstrated by spiking trace amounts of RNA from a fusion positive cell line into circulating RNA from normal research samples, indicating high sensitivity of the detection system. In summary, this study demonstrated the feasibility of gene expression profiling and gene fusion detection from circulating RNA in plasma research samples on Ion Torrent NGS platforms.Fusion Gene Detection and Gene Expression Analysis of Circulating RNA in Plas...

Fusion Gene Detection and Gene Expression Analysis of Circulating RNA in Plas...Thermo Fisher Scientific
Recomendados
The presence of circulating (cell-free) nucleic acids in the bloodstream offers a potential non-invasive approach to monitor disease status and guide treatment options. In past years, increasing interest has been shown for circulating RNA; especially circulating small RNAs for their potential application as biomarkers that may lead toward more effective diagnosis and prognosis in the future. However, widespread inconsistencies have been observed among the studies due to biases generated during sample collection, handling, RNA extraction and analysis. We have developed a complete workflow that includes blood collection, plasma preparation, circulating RNA extraction, followed by expression analysis and gene fusion detection on Ion TorrentTM Next-Generation Sequencing platforms. Blood plasma research samples from normal research samples were utilized for circulating RNA isolation following a TRIzolTM LS Reagent and mirVanaTM miRNA Isolation Kit-based method to maximize circulating RNA recovery. Ion AmpliSeqTM library preparation was performed on purified circulating RNA using either Ion AmpliSeq Transcriptome panel for expression profiling of 21K coding and non-coding genes, or an Ion AmpliSeq panel targeting fusion transcript detection from RNA. Ion AmpliSeq Transcriptome data was analyzed using ampliSeqRNA plugin in Torrent Suite™ Software. ~3000 genes were detected in cfDNA from plasma research samples with high correlation (r>0.8) observed between normal research samples. Ion Reporter™ Software was used to analyze fusion transcript panel data. Detection of fusion gene transcripts was demonstrated by spiking trace amounts of RNA from a fusion positive cell line into circulating RNA from normal research samples, indicating high sensitivity of the detection system. In summary, this study demonstrated the feasibility of gene expression profiling and gene fusion detection from circulating RNA in plasma research samples on Ion Torrent NGS platforms.Fusion Gene Detection and Gene Expression Analysis of Circulating RNA in Plas...

Fusion Gene Detection and Gene Expression Analysis of Circulating RNA in Plas...Thermo Fisher Scientific
Fusion transcripts resulting from translocation events in the oncogenic driver
genes ALK, RET, ROS1, and NTRK1 play an important role in lung
adenocarcinoma. There is a need to detect these fusion transcripts with up to
date technologies as they may serve as viable therapeutic targets. We have
utilized a targeted sequencing approach and developed an Ion AmpliSeq™ RNA
Lung Fusion panel, a workflow, and an Ion Reporter™ analysis solution to detect
these known fusion events. The panel detects transcripts from 37 ALK, 9 RET, 15
ROS1, and 11 NTRK fusion variants along with 5 housekeeping genes to serve
as internal controls. The workflow is FFPE compatible requiring an input of only
10 ng of total RNA with the capacity to multiplex up to 16 libraries on a single Ion
318™ chip. The panel was initially validated using 10ng of total RNA from a
cocktail of 3 cell lines containing known lung cancer fusions (H2228 – EML4-ALK
variant 3a and 3b, HCC78 – SLC34A2-ROS1 and LC-2/ad – CCDC6-RET). The
library was sequenced using the Ion PGM™ system and analyzed with the
AmpliSeq™ RNA Lung Fusion workflow in Ion Reporter™. Analysis showed that
the positive control sample contained all expected fusions and control genes and
reported zero false positives fusions. This multiplexed fusion transcript targeted
sequencing solution is currently being validated by all members of the
OncoNetwork Consortium who will test lung cancer tissue samples that have
been well characterized by FISH, real-time PCR, IHC, and/or massarray. Initial
results from OncoNetwork Consortia members reveal 100% concordance
between the AmpliSeq™ RNA Lung Fusion panel and FISH in 25 lung tissue
samples.Verification of an Ion AmpliSeq™ RNA Fusion Lung Cancer Research Panel, workf...

Verification of an Ion AmpliSeq™ RNA Fusion Lung Cancer Research Panel, workf...Thermo Fisher Scientific
Research has shown that circulating tumor DNA (ctDNA) is
informative of tumor load and tumor evolution in both solid and
hematological cancer. The ability to detect mutations in ctDNA
holds the promise for an accurate and non-invasive approach to
assess minimum residual disease as well as treatment
response in the future. However, as ctDNA often makes up only
a small fraction of cell-free DNA recovered from the plasma,
traditional methods of targeted sequencing research often face
a poor signal-to-noise ratio that cannot be overcome with deep
coverage.
Here we present a novel research method that is capable of
detecting ultra-rare mutations at allelic frequency below 0.5%.
This approach leverages target multiplexing capabilities of the
Ion AmpliSeq™ technology with some important modifications
to the sample preparation procedures. The new protocol
requires as little as 20 ng of input DNA and offers a sample-toanswer
turn-around time in under 24 hours. To support the
analysis of this new approach, we have further developed a
novel Bayesian statistics that models the propagation of
potential artifacts introduced during amplification and sampling
effects during sequencing to differentiate false positives
(variants observed in sequencing data that were not present in
input DNA) from true mutations that were present at very low
levels in the original research sample.
We successfully applied this new method to detect spike-in
mutant DNA in both cell line (Coriell GM24385) and cfDNA
samples. Specifically, we demonstrated the detection of 140
COSMIC genomic aberrations found in 23 frequently mutated
genes. In preliminary study, the method achieved greater 90%
sensitivity and specificity.Successful detection of 40 COSMIC hotspot mutations at allelic frequency belo...

Successful detection of 40 COSMIC hotspot mutations at allelic frequency belo...Thermo Fisher Scientific
We identified mutations in eleven cell free
(cf) DNA samples by next generation
sequencing (NGS) using the Ion AmpliSeq™
Colon & Lung Cancer Research Panel and
the Ion PGM™ System. Since detection of
low frequency mutant alleles may not always
be called confidently in NGS, we verified
results by rare mutation analysis using
digital PCR on the QuantStudio™ 3D Digital
PCR System as an independent method.
We show that frequencies detected are
consistent for both methods for low
frequency mutant alleles at and below 1%.Rare Mutation Analysis Using Digital PCR on QuantStudio™ 3D to Verify Ion Amp...

Rare Mutation Analysis Using Digital PCR on QuantStudio™ 3D to Verify Ion Amp...Thermo Fisher Scientific
The tumor microenvironment (TME) is the intersection between tumor cells and
surrounding non-transformed cells. It contains immune cells, signaling molecules,
stromal and extracellular matrix. Research has shown the TME is often associated
with tumor growth. However, the function and regulatory mechanism of each
constituent is still poorly understood. The presence of PD-L1 is a promising marker
to predict positive response for T cell checkpoint therapy. Current IHC methods to
measure PD-L1 are subjective and highly variable. A higher-throughput and
standardized method that can systematically measure gene expression of cells
present in the TME has emerged to be a more desirable solution.
We applied the OncomineTM Immune Response Research Assay to measure the
expression of 395 genes in non-small cell lung cancer (NSCLC) research samples
from 40 matched FFPE and fresh frozen sample types. This assay covers genes
involved in checkpoint pathway, T cell regulation, cytokine and interferon signaling
pathways, and markers of different tumor infiltrating lymphocyte (TIL) subsets, as
well as tumor markers. With an input requirement of 10 ng of total RNA, libraries
were generated, templated on the Ion ChefTM and sequenced on the Ion S5TM
System. Sequencing data was analyzed and mapped with Torrent Suite Software
and differential expression analysis was conducted with AffymetrixTM Transcriptome
Analysis Console.Gene expression profile of the tumor microenvironment from 40 NSCLC FFPE and ...

Gene expression profile of the tumor microenvironment from 40 NSCLC FFPE and ...Thermo Fisher Scientific
T-Cell receptor (TCR) repertoire sequencing by next-generation
sequencing (NGS) is a valuable tool for building a deeper
understanding of the adaptive immune system. As immunotherapy,
particularly T-cell therapies, show increasing potential in treating
cancer, the ability to gain a detailed, unbiased view of the TCR
repertoire becomes imperative for biomarker discovery, immune
response to treatment, and study of tumor microenvironments. A key
question the field seeks to understand is the relationship between
circulating T-cells and infiltrating T-cells at the tumor site. Here, we
present a novel AmpliSeq approach for TCR repertoire sequencing
using the Ion Torrent S5 sequencer which leverages simplified
workflows and offers up to 600 bp reads which allow for a more
complete characterization of the entire V(D)J region of TCRβ. With a
unique long read length capability, this method can leverage mRNA as
input, which minimizes requirement as starting materials (10-500ng for
typical use cases) and focusing sequencing to productive TCRβ
arrangements.Sequencing the circulating and infiltrating T-cell repertoire on the Ion S5TM

Sequencing the circulating and infiltrating T-cell repertoire on the Ion S5TMThermo Fisher Scientific
The discovery of circulating tumor DNA (ctDNA) in blood, urine
and other bodily fluids has led to a new type of non-invasive
method of characterizing cancer-causing mutations, the liquid
biopsy. With NGS technologies becoming increasingly
sensitive, down to a Limit of Detection (LOD) of 0.1%, they are
rapidly gaining traction as a valid assay for cancer genotyping
and have potential to direct cancer treatment plans. The wideangle
view provided by NGS panels, combined with digital
PCR’s zoomed-in precision detection of DNA provide a
comprehensive picture of a cancer’s genetic makeup. By
applying these complementary techniques at the appropriate
time based on the disease type and stage, cancer treatment
becomes quicker, more precise and more cost-effective in the
future. NGS and digital PCR (dPCR) together provide a
complete picture of the cancer genome.Orthogonal Verification of Oncomine cfDNA Data with Digital PCR Using TaqMan ...

Orthogonal Verification of Oncomine cfDNA Data with Digital PCR Using TaqMan ...Thermo Fisher Scientific
We have developed an amplicon-based NGS approach for FFPE
samples that can detect SNVs, small mutations and LRs
simultaneously. We have implemented a comprehensive
bioinformatics algorithm that detects LRs at high sensitivity, even in
the absence of control sample(s). This significantly reduces the cost
and labor for BRCA1/2 genetic analyses.A next Generation Sequencing Approach to Detect Large Rearrangements in BRCA1...

A next Generation Sequencing Approach to Detect Large Rearrangements in BRCA1...Thermo Fisher Scientific
Mais conteúdo relacionado
Mais procurados
Fusion transcripts resulting from translocation events in the oncogenic driver
genes ALK, RET, ROS1, and NTRK1 play an important role in lung
adenocarcinoma. There is a need to detect these fusion transcripts with up to
date technologies as they may serve as viable therapeutic targets. We have
utilized a targeted sequencing approach and developed an Ion AmpliSeq™ RNA
Lung Fusion panel, a workflow, and an Ion Reporter™ analysis solution to detect
these known fusion events. The panel detects transcripts from 37 ALK, 9 RET, 15
ROS1, and 11 NTRK fusion variants along with 5 housekeeping genes to serve
as internal controls. The workflow is FFPE compatible requiring an input of only
10 ng of total RNA with the capacity to multiplex up to 16 libraries on a single Ion
318™ chip. The panel was initially validated using 10ng of total RNA from a
cocktail of 3 cell lines containing known lung cancer fusions (H2228 – EML4-ALK
variant 3a and 3b, HCC78 – SLC34A2-ROS1 and LC-2/ad – CCDC6-RET). The
library was sequenced using the Ion PGM™ system and analyzed with the
AmpliSeq™ RNA Lung Fusion workflow in Ion Reporter™. Analysis showed that
the positive control sample contained all expected fusions and control genes and
reported zero false positives fusions. This multiplexed fusion transcript targeted
sequencing solution is currently being validated by all members of the
OncoNetwork Consortium who will test lung cancer tissue samples that have
been well characterized by FISH, real-time PCR, IHC, and/or massarray. Initial
results from OncoNetwork Consortia members reveal 100% concordance
between the AmpliSeq™ RNA Lung Fusion panel and FISH in 25 lung tissue
samples.Verification of an Ion AmpliSeq™ RNA Fusion Lung Cancer Research Panel, workf...

Verification of an Ion AmpliSeq™ RNA Fusion Lung Cancer Research Panel, workf...Thermo Fisher Scientific
Research has shown that circulating tumor DNA (ctDNA) is
informative of tumor load and tumor evolution in both solid and
hematological cancer. The ability to detect mutations in ctDNA
holds the promise for an accurate and non-invasive approach to
assess minimum residual disease as well as treatment
response in the future. However, as ctDNA often makes up only
a small fraction of cell-free DNA recovered from the plasma,
traditional methods of targeted sequencing research often face
a poor signal-to-noise ratio that cannot be overcome with deep
coverage.
Here we present a novel research method that is capable of
detecting ultra-rare mutations at allelic frequency below 0.5%.
This approach leverages target multiplexing capabilities of the
Ion AmpliSeq™ technology with some important modifications
to the sample preparation procedures. The new protocol
requires as little as 20 ng of input DNA and offers a sample-toanswer
turn-around time in under 24 hours. To support the
analysis of this new approach, we have further developed a
novel Bayesian statistics that models the propagation of
potential artifacts introduced during amplification and sampling
effects during sequencing to differentiate false positives
(variants observed in sequencing data that were not present in
input DNA) from true mutations that were present at very low
levels in the original research sample.
We successfully applied this new method to detect spike-in
mutant DNA in both cell line (Coriell GM24385) and cfDNA
samples. Specifically, we demonstrated the detection of 140
COSMIC genomic aberrations found in 23 frequently mutated
genes. In preliminary study, the method achieved greater 90%
sensitivity and specificity.Successful detection of 40 COSMIC hotspot mutations at allelic frequency belo...

Successful detection of 40 COSMIC hotspot mutations at allelic frequency belo...Thermo Fisher Scientific
We identified mutations in eleven cell free
(cf) DNA samples by next generation
sequencing (NGS) using the Ion AmpliSeq™
Colon & Lung Cancer Research Panel and
the Ion PGM™ System. Since detection of
low frequency mutant alleles may not always
be called confidently in NGS, we verified
results by rare mutation analysis using
digital PCR on the QuantStudio™ 3D Digital
PCR System as an independent method.
We show that frequencies detected are
consistent for both methods for low
frequency mutant alleles at and below 1%.Rare Mutation Analysis Using Digital PCR on QuantStudio™ 3D to Verify Ion Amp...

Rare Mutation Analysis Using Digital PCR on QuantStudio™ 3D to Verify Ion Amp...Thermo Fisher Scientific
The tumor microenvironment (TME) is the intersection between tumor cells and
surrounding non-transformed cells. It contains immune cells, signaling molecules,
stromal and extracellular matrix. Research has shown the TME is often associated
with tumor growth. However, the function and regulatory mechanism of each
constituent is still poorly understood. The presence of PD-L1 is a promising marker
to predict positive response for T cell checkpoint therapy. Current IHC methods to
measure PD-L1 are subjective and highly variable. A higher-throughput and
standardized method that can systematically measure gene expression of cells
present in the TME has emerged to be a more desirable solution.
We applied the OncomineTM Immune Response Research Assay to measure the
expression of 395 genes in non-small cell lung cancer (NSCLC) research samples
from 40 matched FFPE and fresh frozen sample types. This assay covers genes
involved in checkpoint pathway, T cell regulation, cytokine and interferon signaling
pathways, and markers of different tumor infiltrating lymphocyte (TIL) subsets, as
well as tumor markers. With an input requirement of 10 ng of total RNA, libraries
were generated, templated on the Ion ChefTM and sequenced on the Ion S5TM
System. Sequencing data was analyzed and mapped with Torrent Suite Software
and differential expression analysis was conducted with AffymetrixTM Transcriptome
Analysis Console.Gene expression profile of the tumor microenvironment from 40 NSCLC FFPE and ...

Gene expression profile of the tumor microenvironment from 40 NSCLC FFPE and ...Thermo Fisher Scientific
T-Cell receptor (TCR) repertoire sequencing by next-generation
sequencing (NGS) is a valuable tool for building a deeper
understanding of the adaptive immune system. As immunotherapy,
particularly T-cell therapies, show increasing potential in treating
cancer, the ability to gain a detailed, unbiased view of the TCR
repertoire becomes imperative for biomarker discovery, immune
response to treatment, and study of tumor microenvironments. A key
question the field seeks to understand is the relationship between
circulating T-cells and infiltrating T-cells at the tumor site. Here, we
present a novel AmpliSeq approach for TCR repertoire sequencing
using the Ion Torrent S5 sequencer which leverages simplified
workflows and offers up to 600 bp reads which allow for a more
complete characterization of the entire V(D)J region of TCRβ. With a
unique long read length capability, this method can leverage mRNA as
input, which minimizes requirement as starting materials (10-500ng for
typical use cases) and focusing sequencing to productive TCRβ
arrangements.Sequencing the circulating and infiltrating T-cell repertoire on the Ion S5TM

Sequencing the circulating and infiltrating T-cell repertoire on the Ion S5TMThermo Fisher Scientific
The discovery of circulating tumor DNA (ctDNA) in blood, urine
and other bodily fluids has led to a new type of non-invasive
method of characterizing cancer-causing mutations, the liquid
biopsy. With NGS technologies becoming increasingly
sensitive, down to a Limit of Detection (LOD) of 0.1%, they are
rapidly gaining traction as a valid assay for cancer genotyping
and have potential to direct cancer treatment plans. The wideangle
view provided by NGS panels, combined with digital
PCR’s zoomed-in precision detection of DNA provide a
comprehensive picture of a cancer’s genetic makeup. By
applying these complementary techniques at the appropriate
time based on the disease type and stage, cancer treatment
becomes quicker, more precise and more cost-effective in the
future. NGS and digital PCR (dPCR) together provide a
complete picture of the cancer genome.Orthogonal Verification of Oncomine cfDNA Data with Digital PCR Using TaqMan ...

Orthogonal Verification of Oncomine cfDNA Data with Digital PCR Using TaqMan ...Thermo Fisher Scientific
We have developed an amplicon-based NGS approach for FFPE
samples that can detect SNVs, small mutations and LRs
simultaneously. We have implemented a comprehensive
bioinformatics algorithm that detects LRs at high sensitivity, even in
the absence of control sample(s). This significantly reduces the cost
and labor for BRCA1/2 genetic analyses.A next Generation Sequencing Approach to Detect Large Rearrangements in BRCA1...

A next Generation Sequencing Approach to Detect Large Rearrangements in BRCA1...Thermo Fisher Scientific
Mais procurados (20)
Verification of an Ion AmpliSeq™ RNA Fusion Lung Cancer Research Panel, workf...

Verification of an Ion AmpliSeq™ RNA Fusion Lung Cancer Research Panel, workf...
Automated DNA purification from diverse Microbiome samples using dedicated Mi...

Automated DNA purification from diverse Microbiome samples using dedicated Mi...
Stable 16 year storage of DNA purified with the QIAamp® DNA Blood mini kit - ...

Stable 16 year storage of DNA purified with the QIAamp® DNA Blood mini kit - ...
Use of Methylation Markers for Age Estimation of an unknown Individual based ...

Use of Methylation Markers for Age Estimation of an unknown Individual based ...
Sequential Automation of RNA and DNA preps on the same QIAcube instrument

Sequential Automation of RNA and DNA preps on the same QIAcube instrument
An NGS workflow to detect down to 0.1% allelic frequency in cfDNA

An NGS workflow to detect down to 0.1% allelic frequency in cfDNA
IRJET- Silencing of hnRNP A1 and hnRNP A2/B1 Downregulates the Expression of ...

IRJET- Silencing of hnRNP A1 and hnRNP A2/B1 Downregulates the Expression of ...
Automated DNA extraction from FFPE tissue using a xylene free deparaffinizati...

Automated DNA extraction from FFPE tissue using a xylene free deparaffinizati...
Advances in Breast Tumor Biomarker Discovery Methods

Advances in Breast Tumor Biomarker Discovery Methods
Successful detection of 40 COSMIC hotspot mutations at allelic frequency belo...

Successful detection of 40 COSMIC hotspot mutations at allelic frequency belo...
Rare Mutation Analysis Using Digital PCR on QuantStudio™ 3D to Verify Ion Amp...

Rare Mutation Analysis Using Digital PCR on QuantStudio™ 3D to Verify Ion Amp...
Deeper Insight into Transcriptomes! Download the Flyer

Deeper Insight into Transcriptomes! Download the Flyer
Gene expression profile of the tumor microenvironment from 40 NSCLC FFPE and ...

Gene expression profile of the tumor microenvironment from 40 NSCLC FFPE and ...
suraj_jaladanki_examining_Malaclemys_terrapin_genome_scaffolds

suraj_jaladanki_examining_Malaclemys_terrapin_genome_scaffolds
Multiplex TaqMan Assays for Rare Mutation Analysis Using Digital PCR

Multiplex TaqMan Assays for Rare Mutation Analysis Using Digital PCR
Rapid extraction of high yield, high quality DNA from tissue samples - Downlo...

Rapid extraction of high yield, high quality DNA from tissue samples - Downlo...
Sequencing the circulating and infiltrating T-cell repertoire on the Ion S5TM

Sequencing the circulating and infiltrating T-cell repertoire on the Ion S5TM
Orthogonal Verification of Oncomine cfDNA Data with Digital PCR Using TaqMan ...

Orthogonal Verification of Oncomine cfDNA Data with Digital PCR Using TaqMan ...
A next Generation Sequencing Approach to Detect Large Rearrangements in BRCA1...

A next Generation Sequencing Approach to Detect Large Rearrangements in BRCA1...
Semelhante a KO poster 8:13
Semelhante a KO poster 8:13 (20)
Technical Guide to Qiagen PCR Arrays - Download the Guide

Technical Guide to Qiagen PCR Arrays - Download the Guide
Whole Transcriptome Analysis of Testicular Germ Cell Tumors

Whole Transcriptome Analysis of Testicular Germ Cell Tumors
CENT-1206APP11.15-A_Characterizing RNA Nanoparticles by Analytical Ultracentr...

CENT-1206APP11.15-A_Characterizing RNA Nanoparticles by Analytical Ultracentr...
Sequencing the Human TCRβ Repertoire on the Ion S5TM

Sequencing the Human TCRβ Repertoire on the Ion S5TM
KO poster 8:13
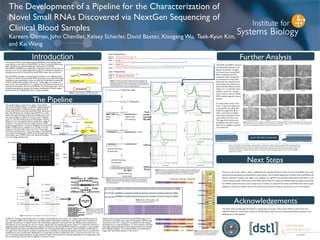
- 1. The Development of a Pipeline for the Characterization of Novel Small RNAs Discovered via NextGen Sequencing of Clinical Blood Samples Kareem Osman, John Chevillet, Kelsey Scherler, David Baxter, Xiaogang Wu, Taek-Kyun Kim, and Kai Wang Introduction The Pipeline Further Analysis Next Steps Acknowledgements Band 1 Band 2 The advent of Next Gen Sequencing and RNAseq has enabled major exploration in the realm of small RNA. Cell-free circulating RNAs have been identified as potential diagnostic biomarkers of disease. MicroRNAs (miRNAs) are especially intriguing, as their function has im- plications on various physiological processes. It is therefore valuable to develop protocols to characterize novel RNAs when they are found. During RNAseq analysis of patient blood samples in an ongoing study of sepsis, hundreds of novel small RNA sequences were disovered. Eight sequences were specifically chosen as they were identified consistently in both whole blood samples and serum. These eight are chosen as a pilot set for the development of a pipeline for the characterization and functional assessment of novel small RNAs.The following project aims to achieve these goals by taking a thorough consideration of each experi- mental factor to ultimately produce a robust pipeline. The pipeline gets its input from either whole blood cell or serum RNA (Figure 2.) cDNAs originating from whole blood RNA are obained from a sepsis study. 8 novel small RNAs previously identified from these sam- ples with RNAseq are verified using qPCR.Two candi- dates with appropriate products by melting curve anal- ysis were chosen to follow, Cad 3 and Cad 7 (Figures 3 and 4.) Gel Electrophoresis and Bioanalyzer analysis (Figure 5.) are next performed to reveal their range of products more specifically.The length of each cDNA products is predicted to result in fragments of minimum length of 80bp (from a sequence of ~20 bp. It is inter- esting to note that both Cad 3 and Cad 7 have larger secondary products form. The next stage in the pipeline is to excise and elute the cDNAs from each gel band to clone them into a TOPO vector for bacterial transfec- tion and Sanger Sequencing. TOPO TA Cloning is next performed to transfect and proliferate Cad 3 and 7 on a larger scale.Transformation of the invitrogen TOPO vector (Figure 6.) into Alpha Select chemically competent E. coli is performed. 6 colonies are selected to proliferate in liquid culture.The recombinant plasmid is extracted with the QiaPrep spin miniprep kit, and aliquots of each sample are sent for Sanger sequencing.The sequencing results (Figure 7.) reveal the variation in fragment length of these RNA products may be functional, as the queried RNA sequence is present in each product. With evidence that these identified small RNAs are real and polymorphic, another round of qPCR is perfomed on the sepsis and control plasma samples, revealing more candidates to develop the pipeline with.There is an observed temperature and concentration dependence in many these products (Figure 8.) Cad 1, Cad 2-5p, and Cad 3 are cho- sen to continue experimentation, as they consist of either one product or the least dependence on temperature and not concentration, factors that are difficult to control in-vitro. Figure 7. Analysis of the Sanger sequencing results reveals that the smallest of the multiple products observed via gel elecrophoresis and Bioanalyzer analysis all contain the query sequence of the RNA product (red), followed by the poly-A tail and Quantitect universal tag (blue), and sometimes followed by an unknown sequence (green). Figure 9. Reverse Transcription of NCI-60 cell line RNA is next performed using the miScript II RT kit (HiSpec Buffer) to identify cell lines with high rel- ative abundance of all Cad 1, Cad 2, and Cad 3 primer products (indicated by low CT during qPCR). Figure 8. Melt Curve analysis of the qPCR of cDNA using each of the eight small RNA primers reveals an annealing temperature dependance in the the Cad 4 prod- uct (above quadrant, highlighted dark green) and a cDNA concentration depen- dance in the observed products of primers 5, 6, and 7 (lower quadrant). Reverse Transcription of NCI-60 cell line RNA (Figure 9.) is performed to broaden the scope of this pipeline. qPCR using the primers for Cad 1, 2-5p, and 3 is performed and relative abundances are compared (Figure 11.) Once specific cell lines with high relative abundance of all three products are chosen, IDT inhibitors (Figure 10.) of these RNAs are purchased to begin their functional analysis in cell culture. Figure 1. Two of the eight novel sequences are cho- sen first to analyze (referred to as Cad 3 and Cad 7) Figure 3. Gel Image of Cad 3 Products There are still further steps to take in establishing this pipeline.While a number of novel small RNAs have been characterized,sequenced,and assessed for conservation, the functional significance of these novel small RNAs still remains unknown. Function may begin to be analyzed via a qPCR of the purified experimental cell RNAs vs the control samples and/or qPCR of the same RNA with primers for these small RNAs potential targets should they be miRNAs (determined by custom target scan). If evidence is obtained that these small RNAs have roles in gene regulation, microarray analysis may then be performed to discover systemic alterations to the transcriptome. We thank those funding agencies below for supporting this project. Also, special thanks to Arian Smit who helped facilitate the conservation study and the ISB Sequencing Core who maintained facilities essential to the development of this pipeline. Figure 4. Gel Image of Cad 7 Products Figure 2. RNA may be extracted from both whole blood and serum samples. cDNAs obtained from whole blood samples provide the initial input to the pipeline cle Figure 5. Bioanalyzer Chromatograph of Cad 4 and Cad 7 products. Figure 6. The invitrogenTOPO TA Cloning kit was used to gen- erate recombinant plasmid for transfection of alpha select K12 strain Ecoli. Figure 10. IDT inhibitor design: a Zen modified reverse and complement strand to each novel small RNA analyzed Figure 11. Heatmap generated to analyze the relative abundance of Cad 1, Cad 2, and Cad 3 products in the NCI-60 and LNCaP,Tera1, 293T cell lines via CT threshold analysis Figure 12. Phase Contrast imaging of transfected and control of A549 and SNB19 cells 24 and 48 hours after transfection with IDT inhibitors.There does not appear to be any indication of morphologic change or reduced cell viability when comparing the cell density of the experimental groups with that of the negative controls. The A549 and SNB19 cell lines are ultimately selected to un- dergo the inhibitor transfec- tion experiment (using RNAi- MAX Lipofectamine).Two timepoints after transfection are examined to empirically determine the required time- frame before the cultures are lysed and their RNAs purified (Figure 12.) A miR-34a mimic positive control for transfec- tion is used, as well as a RNAi- MAX-treated negative control (water). A conservation study of the Cad 1, 2-5p, and 3 sequences is also performed using the PhastCons algorithm. Both Cad 1 and Cad 3 are highly conserved in eutherian mam- mals, while Cad 2-5p is not. The algorithm indicates the probability that each nucleo- tide and its neighbors queried are conserved. Figure 13. The UCSC Genome Browser is used to perform a conservation analysis of Cad 1-3 through a BLAT and subsequent anal- ysis using the PhastCons algorithm. PhastCons indicates that both Cad 1 and Cad 3 have 100% base-pair conservation through ~70% of the query length when compared against the UCSC database of 100 vertebrates.These results furthermore indicate complete conservation in eutherian mammals. Cad 2-5p, however, is not highly conserved, displaying a PhastCons score of 0 across alleach basepair of its sequence. P50 GM076547
