Packaging of DNA helix...
•
5 gostaram•1,522 visualizações
The document discusses several key concepts related to DNA packaging and gene expression. It describes the central dogma where genetic information flows from DNA to RNA to proteins. It also discusses how DNA is packaged in cells through nucleosomes and chromatin, and how chromatin exists in two forms - euchromatin which is loosely packed and transcriptionally active, and heterochromatin which is tightly packed and transcriptionally inactive. It provides information on calculating DNA length and the number of base pairs.
Denunciar
Compartilhar
Denunciar
Compartilhar
Recomendados
Recomendados
Mais conteúdo relacionado
Mais procurados
Mais procurados (20)
Semelhante a Packaging of DNA helix...
Semelhante a Packaging of DNA helix... (20)
Genome organization in eukaryotes (molecular biology)

Genome organization in eukaryotes (molecular biology)
Chromatin Structure & Genome Organization by Shivendra Kumar

Chromatin Structure & Genome Organization by Shivendra Kumar
Chromosomes structure and function, Dr.Kamelsh shah, PSSHDA, KADI 

Chromosomes structure and function, Dr.Kamelsh shah, PSSHDA, KADI
Mais de HARINATHA REDDY ASWARTHA
Symbiotic algae, Measurement of algal growth, Algal strain selection, Cultivation of algae, Biofuel production from algae:Symbiotic algae, Measurement of algal growth, Algal strain selection, Cultiva...

Symbiotic algae, Measurement of algal growth, Algal strain selection, Cultiva...HARINATHA REDDY ASWARTHA
Mais de HARINATHA REDDY ASWARTHA (20)
Structure of proteins and nature of bond linking monomers in a polymer

Structure of proteins and nature of bond linking monomers in a polymer
FOXP2 gene mutated in a speech and language disorder

FOXP2 gene mutated in a speech and language disorder
Mycorrhizae ecto and endo mycorrhizae significance

Mycorrhizae ecto and endo mycorrhizae significance
Symbiotic algae, Measurement of algal growth, Algal strain selection, Cultiva...

Symbiotic algae, Measurement of algal growth, Algal strain selection, Cultiva...
Algae classification features and reproduction of algae 

Algae classification features and reproduction of algae
Último
https://app.box.com/s/x7vf0j7xaxl2hlczxm3ny497y4yto33i80 ĐỀ THI THỬ TUYỂN SINH TIẾNG ANH VÀO 10 SỞ GD – ĐT THÀNH PHỐ HỒ CHÍ MINH NĂ...

80 ĐỀ THI THỬ TUYỂN SINH TIẾNG ANH VÀO 10 SỞ GD – ĐT THÀNH PHỐ HỒ CHÍ MINH NĂ...Nguyen Thanh Tu Collection
Último (20)
80 ĐỀ THI THỬ TUYỂN SINH TIẾNG ANH VÀO 10 SỞ GD – ĐT THÀNH PHỐ HỒ CHÍ MINH NĂ...

80 ĐỀ THI THỬ TUYỂN SINH TIẾNG ANH VÀO 10 SỞ GD – ĐT THÀNH PHỐ HỒ CHÍ MINH NĂ...
HMCS Vancouver Pre-Deployment Brief - May 2024 (Web Version).pptx

HMCS Vancouver Pre-Deployment Brief - May 2024 (Web Version).pptx
Kodo Millet PPT made by Ghanshyam bairwa college of Agriculture kumher bhara...

Kodo Millet PPT made by Ghanshyam bairwa college of Agriculture kumher bhara...
General Principles of Intellectual Property: Concepts of Intellectual Proper...

General Principles of Intellectual Property: Concepts of Intellectual Proper...
Unit 3 Emotional Intelligence and Spiritual Intelligence.pdf

Unit 3 Emotional Intelligence and Spiritual Intelligence.pdf
Jual Obat Aborsi Hongkong ( Asli No.1 ) 085657271886 Obat Penggugur Kandungan...

Jual Obat Aborsi Hongkong ( Asli No.1 ) 085657271886 Obat Penggugur Kandungan...
Fostering Friendships - Enhancing Social Bonds in the Classroom

Fostering Friendships - Enhancing Social Bonds in the Classroom
Unit-V; Pricing (Pharma Marketing Management).pptx

Unit-V; Pricing (Pharma Marketing Management).pptx
Food safety_Challenges food safety laboratories_.pdf

Food safety_Challenges food safety laboratories_.pdf
Python Notes for mca i year students osmania university.docx

Python Notes for mca i year students osmania university.docx
ICT Role in 21st Century Education & its Challenges.pptx

ICT Role in 21st Century Education & its Challenges.pptx
Packaging of DNA helix...
- 1. • Central dogma • Packaging of DNA Helix • Nucleosome • Euchromatin and Heterochromatin..
- 3. Central dogma • Francis Crick proposed the Central dogma. • The flows of genetic information from DNA to RNA to proteins is called Central dogma.
- 4. • In some viruses (RNA viruses) the flow of information is in reverse direction, that is, from RNA to DNA. RNA DNA • Synthesis of RNA form DNA is called Reverse transcription. • Enzyme involved in this process is Reverse transcriptase.. • or RNA dependent DNA Polymerase enzyme.. Reverse transcription
- 5. Packaging of DNA Helix
- 6. • The length of DNA is calculated by multiplying the total number of bp with distance between two consecutive bp. • The length of DNA = Total no.of base pairs × Distance between two base pairs. • For example: 6.6 × 10 9 bp × 0.34 × 10-9 m/bp = 2.2 metres. How to calculate length of DNA ?
- 7. If the length of E. coli DNA is 1.36 mm, can you calculate the number of base pairs in E.coli? – 1.36mm is DNA length = 1.36 × 10-3 M – 1.36 × 10-3 M / 0.34 × 10-9 M – 4.6 × 10 6 base pairs Total no of base pairs= Total length of DNA / Distance between two base pairs
- 8. Prokaryotes DNA • In prokaryotes, such as, E. coli, though they do not have a defined nucleus. • The DNA is not scattered throughout the cell.
- 10. In prokaryotes • DNA (being negatively charged) is held with some proteins (that have positive charges) in a region termed as ‘nucleoid’. • The DNA in nucleoid is organised in large loops held by proteins.
- 11. In eukaryotes • The length of DNA greater than the dimension of a typical nucleus (approximately 10-6 m)… • A set of positively charged, basic proteins called histones involved in packaging.
- 12. • Histones are rich in the basic amino acid residues lysines and arginines. • Both the amino acid residues carry positive charges in their side chains.
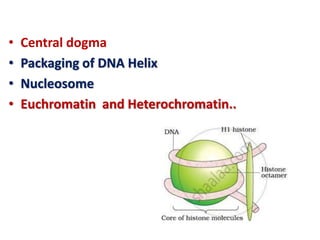
- 14. • Histones are organised to form a unit of eight molecules called as histone octamer. • The negatively charged DNA is wrapped around the positively charged histone octamer to form a structure called nucleosome..
- 16. • A typical nucleosome contains 200 bp of DNA helix. • Nucleosomes constitute the repeating unit of a structure in nucleus called chromatin. •
- 17. • Chromatin thread-like stained (coloured) bodies seen in nucleus. • The nucleosomes in chromatin are seen as ‘beads-on- string’ structure when viewed under electron microscope (EM) .. •
- 18. • The beads-on-string structure in chromatin is packaged to form chromatin fibers.. • Chromatin fibers are further coiled and condensed at metaphase stage of cell division to form chromosomes.
- 20. Non-histone Chromosomal (NHC) proteins. • The packaging of chromatin at higher level requires additional set of proteins that collectively are referred to as Non-histone Chromosomal (NHC) proteins.
- 22. Euchromatin • In a typical nucleus, some region of chromatin are loosely packed (and stains light) and are referred to as euchromatin. • Euchromatin is said to be transcriptionally active chromatin.
- 23. Heterochromatin • The chromatin that is more densely packed and stains dark are called as Heterochromatin. • Heterochromatin is transcriptionally inactive..
- 24. • Shine dalgarno sequence:
- 25. • A sequence of five to nine (typically seven) nucleotides length. • Present before the start codon in prokaryotic messenger RNA (mRNA) that is recognized by the small subunit of ribosome during initiation of translation.
- 26. P and Q arm of chromosome
- 28. Punctuation codons • There are two punctuation marks in the genetic code called the START and STOP codons ..
- 29. No. of nucleosome in Diploid human cell • A diploid human cell contain approximately 30 million nucleosomes..
- 30. Dr. HarinathaReddy Aswartha Assistant professor Department of Microbiology ANDHRAPRADESH INDIA
