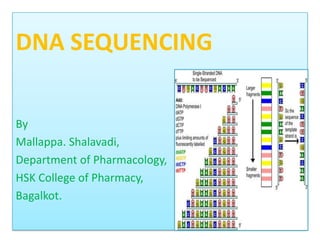
Dna sequencing
- 1. DNA SEQUENCING By Mallappa. Shalavadi, Department of Pharmacology, HSK College of Pharmacy, Bagalkot.
- 2. Contents: 1. Introduction. 2. Methods of sequencing. Conventional DNA sequencing method. Cycle sequencing. Automated DNA sequencing. Pyrosequencing.
- 3. INTRODUCTION: The information content of DNA is encoded in the form of four bases (A,G,C and T) and the process of determining sequence of these bases in a given DNA molecule is referred to as DNA sequencing. DNA fragments can be analyzed to determine the nucleotide sequence of DNA and to determine the distribution and location of restriction sites.
- 4. Fundamental reasons for knowing the sequence of DNA molecule: To charecterise the newly cloned DNA. For predictions about its fuctions. To facilitate manipulation of the molecule. To confirm the identity of a clone or a mutation. To check the fidility of newly created mutation and ligation junction. Screening tool to identify polymorphisms and mutation in genes of particular interest.
- 5. To confirm the product of a PCR. METHODS OF DNA SEQUENCING: (A)Conventional DNA sequencing methods 1. Chemical degradation method. 2. Chain termination method. (B) Cycle sequencing. (C) Automated DNA sequencing. (D) Pyrosequencing.
- 6. (A)Conventional DNA sequencing methods: 1. Chemical degradation method [ Maxam and Gilbert’s method] • This method involves the base specific chemical cleavage of an end labeled DNA segment to generates a set of labeled molecules. PRINCIPLE: • The partially cleaved DNA fragment is subjected to five separate chemical reactions. • Each of which is specific for a particular base.
- 7. • The resulting fragment terminate at that specific base followed by high resolution gel electrophoresis and detection of the labeled fragments by autoradiography METHOD • Labeled DNA at one end with 32P. • DNA copies are divided in to 4 samples. • Each samples treated with a chemical that specifically destroys one or two of the 4 base in DNA.
- 8. Different types of base specific reactions used in the Chemical degradation method. Reagents Base Specific modification Dimethyl Sulphate G Methylation of N7 renders the C8-C9 (pH) bond susceptible to cleavage. Piperidine formate A+G Weakens the glycosidic bond of adenine and guanine residues by protonaing nitrogen atoms in the purin rings resulting in depurination. Hydrazine C+T Opens pyrimidine rings, which recyclize in a five membered form which is valnerable. Hydrazine + 1.5 M Nacl C Only cytocin reacts with Hydrazine.
- 9. • Results in series of labeled fragments. • Length is depends on the distance of distroyed base from the labeled end of molecule. • If G is 3,6 and 9 base away from the labeled end then treatment of DNA strand with chemical that cleave at G will generates labeled fragments 2, 5 and 8 base length. • Acrylamide gel electrophoresis. • Gel is autoradiographed. • Read out sequence.
- 10. • T • G • A • C • G • C • T • G • A • T
- 11. 2. Chain Termination Method [Sanger’s Dideoxy method] • Developed by Frederic’s Sanger. • Common method. • Involves controlled synthesis of DNA to generate fragments terminating at specific point. PRINCIPLE • Replacement of dNTPs with 2’, 3’ dideoxy NTPs in the DNA chain terminates DNA synthesis.
- 12. • This is because these ddNTPs are nucleotide analogues that lacks the 3’ OH group that is necessary for phosphodiester bond formation and chain elongation.
- 13. METHOD • Primer is labeled so that newely synthesised DNA can be detected. • When the primer extended DNA polymerase occasionally inserts a ddNTPs instead of dNTPs. • No further elongation. • In ddA reaction all cains ends with ddA. • DNA chains in in each reaction are seperated by polyacrylamide gel electrophoresis. • Sequence are determined by autoradiogram by reading sequencing ladder from bottom to top to give the sequence in 5’- 3’ orientation.
- 14. (B) Cycle sequencing • Dideoxy mediated sequencing reactions using PCR and end labeled primers. • Also called thermal DNA sequencing or linear amplification of DNA sequencing. • Involves heating reaction mixture to 940 C to denature the template. • Cooling below the melting temperature of primer to allow annealing and repeating the sequencing reaction. • This procedure can be repeated untill one of reaction components is exhausted.
- 15. • 4 seperate amlification reactions are set up . • Each having the same primer and different ddNTP. • 2 cycling programs are used in cycle sequencing.
- 16. I- program • Reaction mixtures are subjected to 15-40 cycles of conventional PCR cycling - Determination of the ds DNA strand. - Annealing of a labeled sequencing primer to its target sequence. - Extention of the anneal primer by a thermostable DNA polymerase. • Finally termination of extended strand is done by the incorporation of ddNTP. • Results in double stranded hybrid.
- 17. • This hybrid is denatured during the first step of the next cycle there by liberating the template strand for another round of priming extension and termination. • Therefore the radio labeled chain terminated products accumulate in a linear fashion. II- program • Annealing step is eliminated so no further extension of primer is possible. • Radio labeled products are finally resolved on a polyacrylamide gel and visualized by autoradiography.
- 18. (C) Automated DNA sequencing • Key advantage is automated data collection in an easy way and in lesser time. • Florescence technique is used for detection of DNA bands. Types of florescence labeling systems 1. FOUR REACTION/ ONE GE SYSTEM • Fluorescent primer are used with non labeled ddNTPs. • Different fluorescence dye are used for 4 chain exyention reaction.
- 19. • Resulting DNA strands are seperated in 4 different lane in electrophoresis. • Detected by fluorescence detector.
- 20. 2. ONE REACTION/ ONE GEL SYSTEM • 4 ddNTPs are labeled with different fluorophor. • Chain extention reaction is carried in a single tube. • Resulting fragment is subjected to gel electrophoresis in a single lane. • The tag is incorporated in DNA molecule. • Leads to termination and attachment of fluorophor at the end of DNA molecules. • Gel is illuminated with argon beam snd detected by photomultiplyer.
- 23. (D) Pyrosequencing • More rapid minisequqncing method. • Not require electrophoresis or any other fragment seperation. • Determine which of the 4 bases is incorporate at each step in the copying of DNA templete. • ddNTPs are not required. • As the new strad is being made, the order in which the dNTPs are incorporated is detected. • So the sequence can read as the reaction proceeds.
- 24. • In reaction all 4 dNTPs are not added at one time. • Each dNTP is added individually in a sequential manner. • If perticular dNTP are not incorporated then it is rapidly degraded by nucleotidase or by washed before addition of next dNTP. • Incorporation leads release of pyrophosphate which is detected in an enzyme cascade that emits light.
- 26. TYPES OF PYROSEQUENCING 1. SOLID PHASE SEQUENCING • Template and primer are immobilized on solid support. • All dNTPs are added stepwise and incorporation of particular dNTP is detected by addition of ATP sulfurylase and luciferase. • A washing step is carried out after addition of each dNTP for removal of the excess sustrates.
- 28. REFERANCE 1. Genetic Engineering by Smitha Rasthogi and N. Pathak. 2. I Genetics 2nd ed by Peter J. Russell
- 29. 2. LIQUID PHASE PYROSEQUEN CING • All reagents with DNA template are added in well of a micro titer plate. • Further steps similar to solid phase sequencing.
- 30. Referance 1. Genetics Engineering by S. Rasthogi and N. Pathak. 2. I Genetics 2nd ed by Peter J. Russell. 3. www.google. Com
